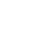
Cardiac Comprehensive Kit analyzes 292 genes and covers major inherited cardiovascular disorders.
Genes Tested
AARS2, ABCA1, ABCC6, ABCC9, ABCG5, ABCG8, ACAD9, ACADVL, ACTA1, ACTA2, ACTC1, ACTN2, ACVR1, ACVR2B, ACVRL1, ADAMTS2, AFF4, AGK, AKAP9, AKT3, ALDH18A1, ALMS1, ALPK3, ANK2, ANKRD1, APOA5, APOB, APOE, ATP6V0A2, ATP6V1A, ATP6V1E1, B3GALT6, B4GALT7, BAG3, BGN, BMPR1B, BMPR2, BRAF, C1R, C1S, CACNA1C, CACNA2D1, CACNB2, CAD, CALM1, CALM2, CALM3, CASQ2, CAV1, CAV3, CBL, CBS, CFAP53, CCND2, CDK13, CELSR1, CELSR2, CELSR3, CHD4, CHD7, CHST14, CITED2, COA5, COA6, COL1A1, COL1A2, COL3A1, COL4A1, COL5A1, COL5A2, COX15, CPT2, CREBBP, CRELD1, CRYAB, CSRP3, CTNNA3, DES, DMD, DNAH11, DNAH5, DNAH6, DNAI1, DNAJC19, DOLK, DSC2, DSE, DSG2, DSP, DTNA, EFEMP2, EHMT1, EIF2AK4, ELAC2, ELN, EMD, ENG, EPHB4, EVC, EVC2, FBLN5, FBN1, FBN2, FHL1, FKBP14, FKTN, FLNA, FLNC, FOXC1, FOXE3, FOXH1, GAA, GANAB, GATA4, GATA5, GATA6, GATAD1, GBE1, GDF1, GJA5, GLA, GNAI2, GORAB, GPC3, GPD1L, GTPBP3, HADHA, HADHB, HAND1, HAND2, HCN4, HRAS, ILK, JAG1, JPH2, JUP, KARS1, KCNA5, KCNAB2, KCND3, KCNE1, KCNE2, KCNE3, KCNH2, KCNJ2, KCNJ5, KCNJ8, KCNK3, KCNQ1, KDM5B, KMT2D, KRAS, LAMA4, LAMP2, LDB3, LDLR, LDLRAP1, LIPA, LMNA, LOX, LPL, LTBP4, LZTR1, MAP2K1, MAP2K2, MED13L, MFAP5, MIB1, MMP21, MRAS, MRPL3, MRPL44, MTO1, MYBPC3, MYH11, MYH6, MYH7, MYL2, MYL3, MYL4, MYLK, MYLK2, MYOZ2, MYPN, NDUFB11, NDUFV2, NEBL, NEXN, NF1, NF2, NIPBL, NKX2-5, NKX2-6, NME7, NODAL, NOTCH1, NOTCH2, NPPA, NR2F2, NRAS, PCSK9, PDLIM3, PIK3CA, PIK3R2, PITX2, PKD1L1, PKP2, PLD1, PLN, PLOD1, POGZ, PPA2, PPP1CB, PRDM16, PRDM5, PRDM6, PRKAG2, PRKD1, PTPN11, PYCR1, RABGAP1L, RAF1, RASA1, RASA2, RBFOX2, RBM10, RBM20, RIT1, RRAS, RYR2, SALL4, SASH1, SCN10A, SCN1B, SCN2B, SCN3B, SCN4B, SCN5A, SCO2, SEMA3D, SEMA3E, SGCD, SHOC2, SKI, SLC22A5, SLC25A20, SLC25A3, SLC25A4, SLC2A10, SLC39A13, SMAD3, SMAD4, SMAD6, SMAD9, SMARCB1, SNTA1, SOS1, SOS2, SPRED1, STAMBP, TAB2, TAFAZZIN, TBX1, TBX20, TBX4, TBX5, TCAP, TECRL, TFAP2B, TGFB2, TGFB3, TGFBR1, TGFBR2, TK2, TLL1, TMEM260, TMEM43, TMEM70, TNNC1, TNNI3, TNNT2, TPM1, TRDN, TRPM4, TSFM, TTN, TTR, VCL, ZEB2, ZFPM2, ZIC3, ZNF469
Technical Specifications
● Genomic regions not covered by this test:
• NM_001374258.1 (BRAF): exon 10 and exon 20
• NM_001042492.3 (NF1): exon 9, 10, 11, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27,28, 29, 32, 33, 34, 35 and 36
• NM_005027.4 (PIK3R2): exon 6
• NM_000384.3 (APOB): exon 1
• NM_015627.3 (LDLRAP1): exon 1
• NM_000237.3 (LPL): exon 5
• NM_014244.5 (ADAMTS2): exon 1
• NM_000722.4 (CACNA2D): exon 14
• NM_000071.3 (CBS): exon 15
• NM_000089.4 (COL1A2): exon 1
• NM_000093.5/NM_001278074.1 (COL5A1): exon 1
• NM_024757.5 (EHMT1): exon 1
• NM_153717.3 (EVC): exon 1
• NM_017617.5 (NOTCH1): exon 1
• NM_016203.4 (PRKAG2): exon 13
• NM_001037.5 (SCN1B): exon 1
• NM_004612.4 (TGFBR1): exon 1
• NM_006073.4 (TRDN): exon 5, 26, and 18
• NM_001267550.2 (TTN): exon 149, 164, 177, 186, 195
• NM_012082.4 (ZFPM2): exon 1
Technical Limitations
The test aims to detect all variants relevant to the genes listed above by targeting all coding exons, of MANE or/and Canonical transcripts, and 12 bp of adjacent intronic sequence. Variants that fall outside of the targeted regions are not intended to be detected by this assay. Unless otherwise noted, sequence changes (SNVs and INDELS) in the promoter and other non-coding regions are not covered by this assay. Certain sequence changes (SNVs and INDELS) in non-coding regions of selected genes that are of clinical significance are also included in the analysis. In cases where two variants are identified in a gene, the test does not distinguish whether these are on one chromosome (in cis) or on different chromosomes (in trans). Certain types of genetic abnormalities such as inversions, rearrangements, polyploidy and epigenetic effects are not covered by this test. Certain sequence changes (SNVs and INDELS) in targeted regions containing repeats, sequences of high homology such as segmental duplications and pseudogenes, as well as regions of high/low GC-content may not be detected. Copy Number Variations (CNVs) are calculated using high quality, de-duplicated and uniquely aligned sequencing reads. CNVs are detected for a subset of the targeted regions using a depth of sequencing coverage approach by applying GC-content normalization. Genomic regions are called as variants if their normalized depth of coverage deviates significantly from the expected normalized coverage which is estimated from a set of reference clinical samples. The test can detect CNVs down to a few exons level resolution. The test cannot detect CNVs at genomic regions with either low mappability or containing repeats, pseudogenes and high/low GC-content. Detection of CNVs using NGS has lower sensitivity/specificity than orthogonal quantification methods, therefore the absence of reported CNVs does not guarantee the absence of CNVs. The lack of disease-causing variants in the targeted genes diminishes but does not exclude the possibility of a disease associated syndrome. Although the test is highly accurate there is still a possibility for false positive or false negative results.