OVERVIEW
DEFINEGlobal developmental delay and intellectual disability (GDD/ID) affect up to 3% of children <5 years old and is defined as a delay in ≥2 developmental domains*. Up to 40% of GDD/ID cases are caused by genetic factors and can occur in isolation or accompanied by other symptoms including malformations and neurological disorders. Children with GDD/ID exhibit mixed and diverse symptoms, and up to two-thirds do not have a single group of symptoms that can point towards a specific diagnosis. As a result, many patients undergo a long diagnostic journey before necessary genetic tests are performed to define the cause of GDD/ID.
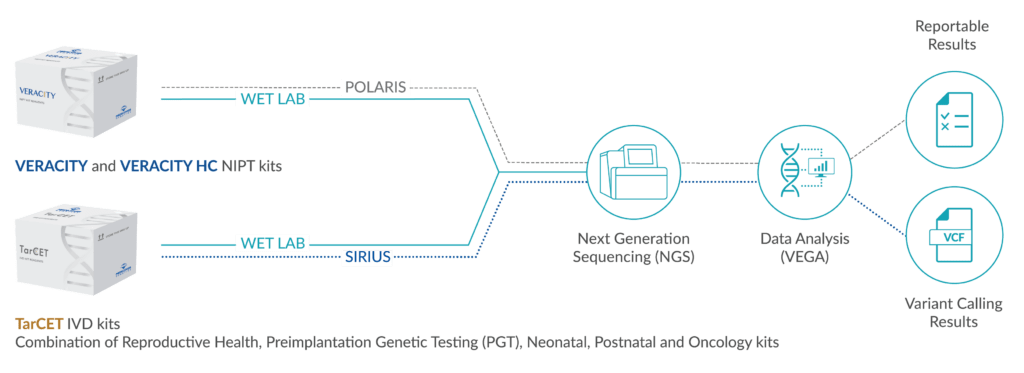
Our tests combine chromosomal analyses and (comprehensive) gene panels associated with many different disorders with overlapping features, providing a diagnostic solution for children with GDD/ID.
Having a diagnosis can help you decide on a management plan or treatment options for your child. Our genetic counselling offers information regarding the diagnosis, identifies associated medical risks, and provides a long-term prognosis, thereby improving your child’s clinical outcome and may help prevent further complications.
*Developmental domains include physical, cognitive, speech/language, social and emotional

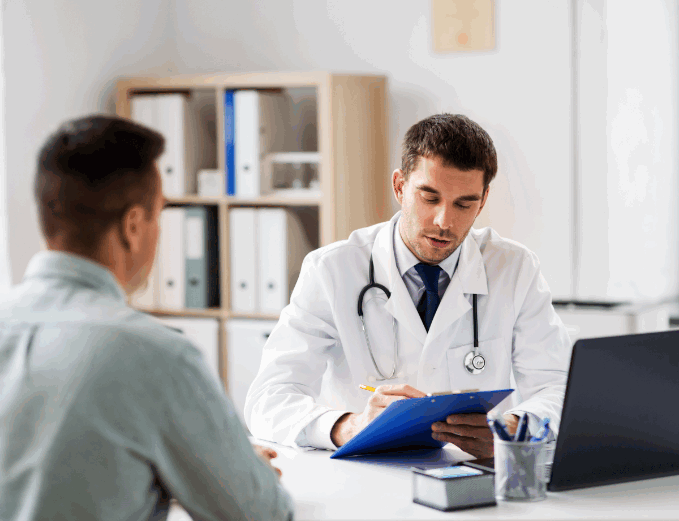
IMPORTANCE OF GETTING TESTED
Children with a GDD/ID disorder often require lifelong support, which can have a profound effect on their lives and that of their family. A timely diagnosis is crucial for therapeutic intervention and the best outcome for your child. Identifying the cause can provide a prognosis, refine treatment options, evaluate recurrence risks and provide closure to the diagnostic journey. In turn, this can improve your mental health and that of your child, while validating your concerns and empowering you to advocate for your child.

Children <5 years with a
significant delay in
≥2 developmental domains*

Children with an
autism spectrum disorder

Children with dysmorphic features
FRAGILE X
Fragile X syndrome is the most common monogenic inherited cause of intellectual disability. Males are more frequently affected than females, and generally with greater severity.
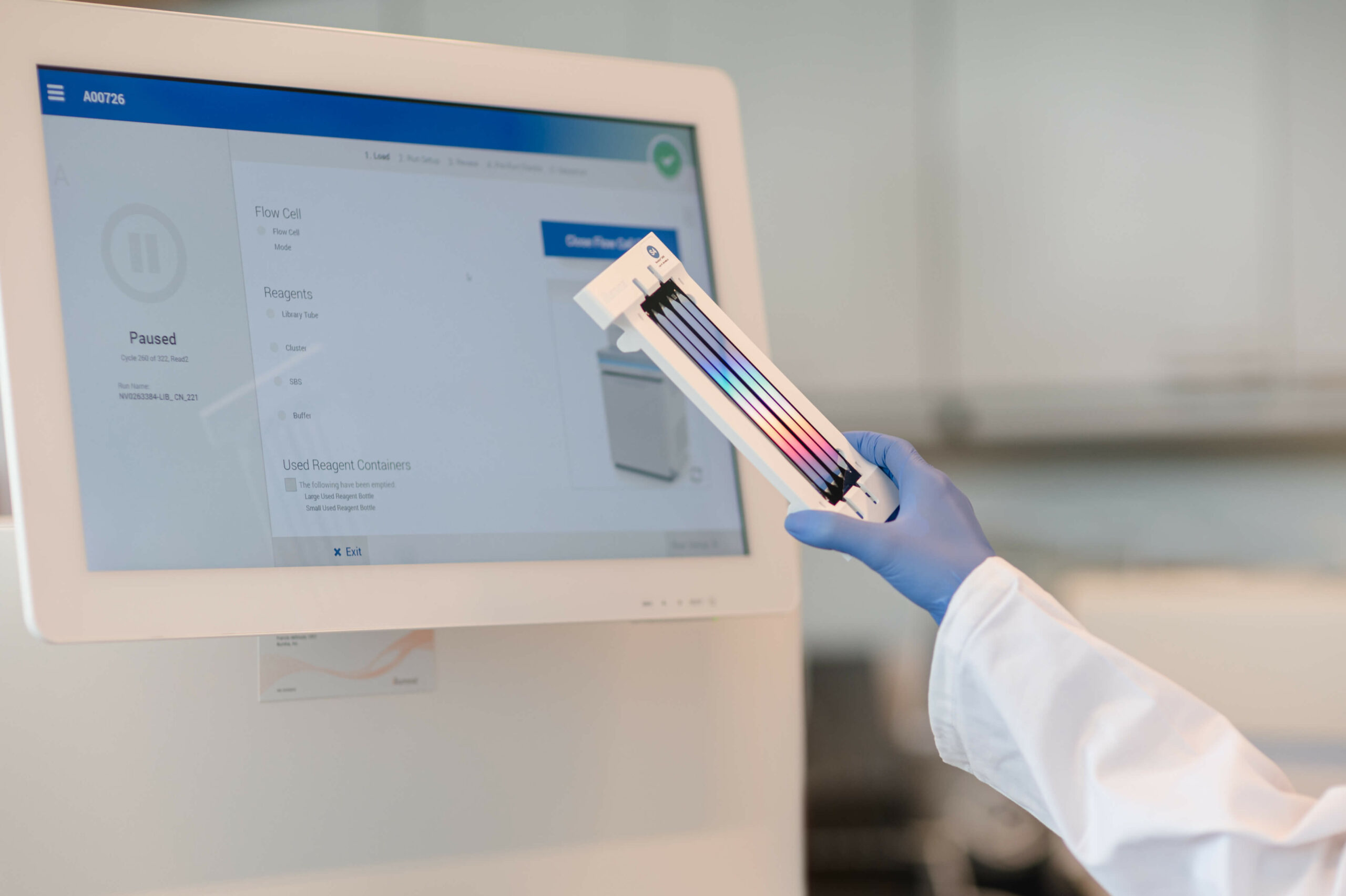
CHROMOSOMAL ANALYSIS
• Karyotyping: detection of gross genetic changes involving several megabases including aneuploidies
• Microarray CGH: screening of deletions and duplications
GENE PANELS
• Autism Spectrum • Brain Malformations (Comprehensive, Lissencephaly, Pontocerebellar Hypoplasia, Tubulinopathies) • Coffin-Siris • Congenital Disorders Of Glycosylation • Cornelia De Lange • GPI Anchor Deficiency • Intellectual Disability • Macrocephaly • Microcephaly • Overgrowth • Pediatric Neurotransmitter • RASopathies, Comprehensive • Rett • Rett And Related Disorders


POSSIBLE OUTCOMES OF THE TEST
A molecular genetic diagnostic report outlining the results of the sequencing analysis is provided. Changes in DNA sequences (variants) can be detrimental and lead to the development of a cardiac or aortic disorder, including asymptomatic disorders that develop later in life. We will report pathogenic and likely pathogenic variants as well as variants of unknown significance.
Pathogenic and likely pathogenic variants mean the genetic cause of the observed symptoms has been identified and may help determine the right treatment and management plan.
Variants of unknown significance means there was not enough evidence to classify the variant as either pathogenic or neutral. Annual variant reclassification and testing family members is recommended.
It is important to note that a negative result does not guarantee the absence of a disorder or that the disorder does not have a genetic cause. Genetic testing is an evolving field and may not detect all variants or there may not currently be enough evidence to classify all variants that lead to an inherited disease.